Summary
In the laboratory, we use different yeasts 1) as an experimental model to study the molecular basis of human congenital glycosylation diseases, focusing on the mechanisms of protein N-glycosylation involved, and 2) as a platform for biotechnological processes: production of recombinant proteins and other metabolites of interest in the pharmacological industry, in bioremediation, and bioenergetics.
Lines of Research
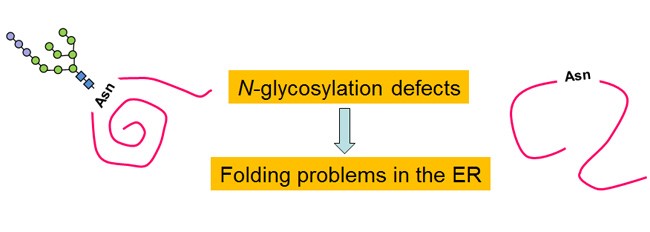
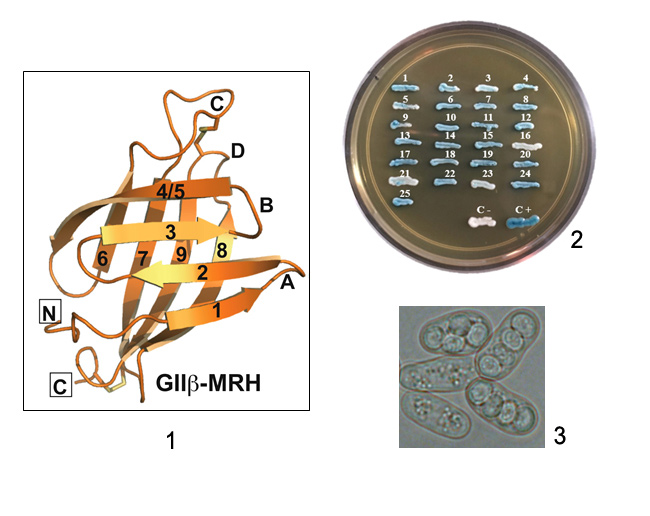
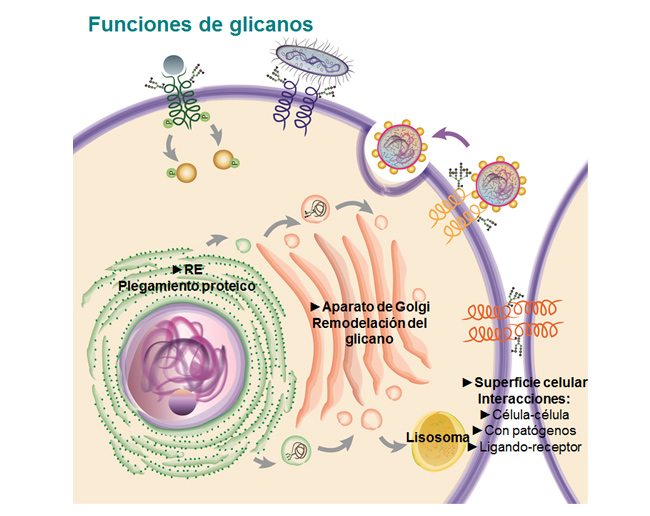
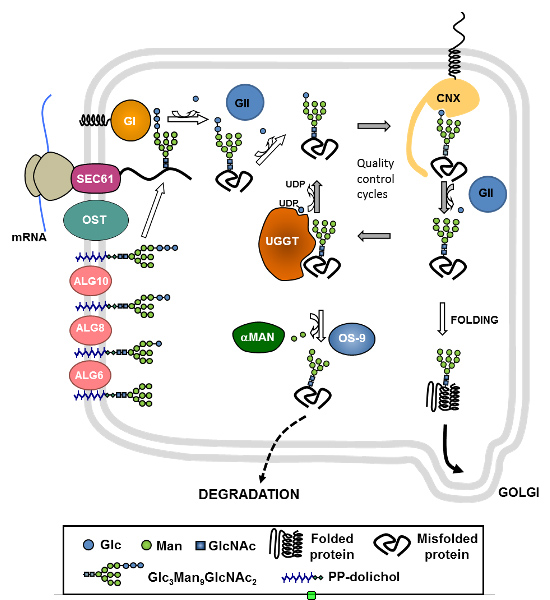
More than 80 human inherited diseases due to defects in protein N-glycosylation are currently known. Type I CDGs are due to defects during the transfer of glycans to proteins, and type II CDGs are due to defects in the subsequent remodeling of glycans. Patients present with multisystem failures appearing at various ages. CDG IIb or MOGS-CDG are caused by mutations in glucosidase I (GI), an ER membrane protein responsible for the first step of N-glycan processing. In the laboratory, we studied the molecular basis of the disease using S. pombe yeast mutants that mimic these defects and explored the cellular causes of the defects produced by the lack of GI, searched for additional functions of this enzyme, and evaluated possible treatments of CDG type IIb.
Yeasts are widely used in the food industry, as a model for the study of eukaryotic cells and as a platform for the expression of recombinant proteins. They are unicellular fungi, many of them non-pathogenic, and with a large number of tools available for biochemical studies and genetic manipulations. In the laboratory, we designed Saccharomyces cerevisiae yeast as a platform for bioremediation processes of dairy industry wastes coupled to bioethanol production, and also as an organism for heterologous production of a photoprotective compound. We also use the yeast Pichia pastoris as a platform for the recombinant production of proteins of biotechnological interest, such as the RBD domain of Sars-CoV2 for use in diagnostic kits or vaccine candidates, or nanoantibodies to be used in replacement of secondary antibodies, among others.
Members:
Sofía Orioli
sorioli.97@gmail.com
Bachelor in Biotechnology | PhD Fellow, CONICET
Victoria Albeck
victoriaalbeck@gmail.com
UBA Fellow | Bachelor Thesis Student (Biology)
Fiorella Iannello Vitale
fioreiannello@hotmail.com
Thesis Student (Industrial Engineering)
LinkedIn
Outreach notes:
Oficina de Vinculación Tecnológica de la FCEN, UBA. Transmitido por YouTube (2021)
Exactas y el Covid-19: Innovación y transferencia
Programa televisivo “Noche de Mente”, TV Publica, 2023
Biotecnología de levaduras
Radio Fénix, FM 100.3, Ciudad de Buenos Aires
Desarrollo de vacuna Argenvac
NexCiencia – Noticias de Ciencia y Técnica
Avance para frenar la pandemia- Proteína buena, barata y abundante
Collaborators:
Dr. Guillermo Alonso
INGEBI
Dr. Nancy Dahms
Medical College Wisconsin
Dr. Diego Libkind
Universidad del Comahue
Dr. Itatí Ibañez
INQUIMAE
Outstanding publications:
- Idrovo-Hidalgo T, Pignataro MF, Bredeston L, Elias F, Herrera MG, Pavan MF, Foscaldi S, Suireszcz M, Fernández NB, Wetzler DE, Paván CH, Craig PO, Roman EA, Ruberto LAM, Noseda DG, Ibañez LI, Czibener, C, Ugalde, JE, Argentinian AntiCovid Consortium, Nadra AD, Santos J, D’Alessio C (2024)
Deglycosylated RBD produced in Pichia pastoris as a low-cost sera COVID-19 diagnosis tool and a vaccine candidate
Glycobiology 34, cwad089
DOI: 10.1093/glycob/cwad089 - Gallo G, Valko A, Herrera Aguilar, N, Weisz, AD y D’Alessio, C (2022)
A novel fission yeast platform to model N-glycosylation and the bases of congenital disorder of glycosylation Type I
Journal of Cell Science
DOI: 10.21769/BioProtoc.4508 - Argentinian AntiCovid Consortium (2022): Berguer PM, Blaustein M, Bredeston LM, Craig PO, D’Alessio C, Elias F, Farré P, Fernández NB, Gentili HG, Gándola YB, Gasulla J, Gudesblat, GE, Herrera MG, Ibañez LI, Idrovo-Hidalgo T, Nadra AD, Noseda DG, Paván CH, Pavan MF, Pignataro MF, Roman EA, Ruberto LAM, Rubinstein N, Sanchez MV, Santos J, Wetzler DE, Zelada AM
Covalent coupling of Spike’s receptor binding domain to a multimeric carrier produces a high immune response against SARS-CoV-2
Scientific Reports 12, 692
DOI: 10.1038/s41598-021-03675-0 - Argentinian AntiCovid Consortium (2020): Arbeitman CR, Auge G, Blaustein M, Bredeston LM, Corapi ES, Craig PO, Cossio LA, Dain L, D’Alessio C, Elias F, Fernandez NB, Gasulla J, Gorojovsky N, Gudesblat GE, Herrera MG, Ibañez LI, Idrovo-Hidalgo T, Iglesias Rando M, Kamenetzky L, Nadra AD, Noseda DG, Pavan CH, Pavan MF, Pignataro MF, Roman EA, Ruberto LAM, Rubinstein N, Santos J, Velazquez F, and Zelada AM
Structural and Functional Comparison of SARS-CoV-2-Spike Receptor Binding Domain Produced in Pichia pastoris and Mammalian Cells
Scientific Reports 10, 21779
DOI: 10.1038/s41598-020-78711-6 - Gallo, GL; Valko, A; Aramburu, SI; Etchegaray, E; Völker, C; Parodi, AJ y D’Alessio, C (2018)
Abrogation of glucosidase I-mediated glycoprotein deglucosylation results in a sick phenotype in fission yeasts: Model for the human MOGS-CDG disorder
Journal of Biological Chemistry (293) 19957- 19973
DOI: 10.1074/jbc.RA118.004844 - Bredeston, LM; Marino-Buslje, C; Mattera, VS; Buzzi, LI; Parodi AJ y D’Alessio C (2017)
The conundrum of UDP-Glc entrance into the yeast ER lumen
Glycobiology 27, 64-79
DOI: 10.1093/glycob/cww092 - D’Alessio, C y Dahms, NM (2015)
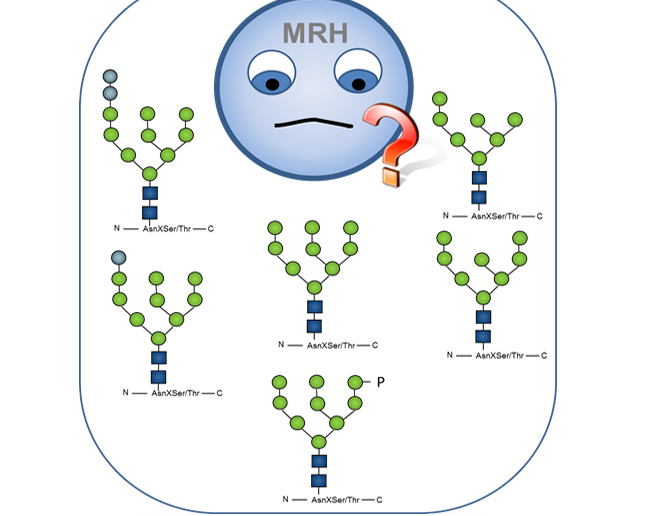
Glucosidase II and MRH-domain containing proteins in the secretory pathway
Current Protein & Peptide Science. 16(1):31-48
DOI: 10.2174/1389203716666150213160438 - Olson, LJ; Orsi, R; Alculumbre, SG; Peterson, FC; Stigliano, ID; Parodi, AJ; D’Alessio, C y Dahms, NM (2013)
Structure of the Lectin MRH Domain of Glucosidase II, an Enzyme that Regulates Glycoprotein Folding Quality Control in the Endoplasmic Reticulum
Journal of Biological Chemistry 288(23):16460-75
DOI: 10.1074/jbc.M113.450239 - Stigliano, ID; Alculumbre, SG; Labriola, CA; Parodi, AJ y D’Alessio, C (2011)
Glucosidase II and N-glycan mannose content regulate the half-lives of monoglucosylated species in vivo
Molecular Biology of the Cell 22, 1810-1823
DOI: 10.1091/mbc.e11-01-0019 - C D’Alessio, JJ Caramelo, AJ Parodi (2010)
UDP-Glc:glycoprotein glucosyltransferase-glucosidase II, the ying-yang of the ER quality control
Semin Cell Dev Biol. 21, 491-9
DOI: 10.1016/j.semcdb.2009.12.014 - Stigliano, ID; Caramelo, JJ; Labriola, CA; Parodi, AJ y D’Alessio, C (2009)
Glucosidase II beta subunit modulates N-glycan trimming in fission yeast and mammals
Molecular Biology of the Cell 20(17):3974-84
DOI: 10.1091/mbc.e09-04-0316 - Olson, LJ; Orsi, R; Alculumbre, SG; Peterson, FC; Stigliano, ID; Parodi, AJ; D’Alessio, C y Dahms, NM (2013)
Structure of the Lectin MRH Domain of Glucosidase II, an Enzyme that Regulates Glycoprotein Folding Quality Control in the Endoplasmic Reticulum
Journal of Biological Chemistry 288(23):16460-75
DOI: 10.1074/jbc.M113.450239
Publications can be found at:
https://scholar.google.com.ar/citations?user=E-p3olYAAAAJ&hl=es